Sfp1 has traditionally been regarded as a ‘classical’ TF known to bind specific promoters, either through interactions with other TFs or by directly binding to promoter DNA (Albert et al., 2019; Reja et al., 2015). Further analysis that we performed using published ChIP-exo and ChEC-seq data has revealed an additional aspect of Sfp1’s binding behavior—it also interacts with gene bodies. Notably, the binding of Sfp1 to CRAC + gene bodies correlates with the presence of transcriptionally active Pol II molecules (Figure 5B), suggesting a physical association between Sfp1 and Pol II. Consequently, the binding of Sfp1 has an impact on Pol II configuration, stoichiometry, and backtracking, particularly in the context of CRAC + genes (Figures 6 and 7). This alteration in the Pol II configuration is reflected in the increased backtracking frequency, as indicated by the BI values. Moreover, the binding of Sfp1 to gene bodies correlates with Pol II activity, particularly within CRAC + genes (Figure 5B). Thus, Sfp1’s function in a subset of its target genes extends beyond transcription initiation and encompasses transcription elongation, as we proposed earlier (Gómez-Herreros et al., 2012b). Based on the following observations, we propose that Sfp1 binds to Pol II in proximity to both DNA and Rpb4, accompanying Pol II during elongation and influencing its configuration, thereby enhancing its propensity to backtrack (Figure 8A): (I) Sfp1 physically interacts with Rpb4, either directly or indirectly (Figure 1—figure supplement 1). (II) ChIP-exo results, dependent on Sfp1’s cross-linking with DNA, indicate Sfp1’s presence near DNA, both at promoters and gene bodies (Figure 5A). (III) Sfp1 influences Pol II configuration in a manner that impacts the architecture or stoichiometry of Rpb4 within Pol II (Figure 7). (IV) Sfp1 binding to chromatin better correlates with all Pol II (both backtracked and active) than with transcriptionally active Pol II, examined by BioGRO-seq (Figure 5B). (V) The global reduction in BI is observed upon the deletion of SFP1 (Figures 6 and 7).
(A) Two modes of action of Sfp1 in transcription: CRAC + genes recruit Sfp1 to their promoters, whereas non-CRAC + genes recruit Sfp1 from the nuclear space directly to RNA polymerase II (Pol II). Upper panel represents CRAC + genes that recruit Sfp1 to their promoters. We discovered that Sfp1 appears to accompany Pol II of CRAC + genes, in a manner proportional to the number of transcriptionally active Pol II (Figure 5A–B). Its binding to all Pol II molecules, including backtracked Pol II, is even more apparent (Figure 5B). In addition, we found that CRAC + genes are enriched with Rap1-binding sites (Figure 3). We, therefore, propose that, following binding to Rap1-containing promoters, Sfp1 binds Pol II. Specifically, it binds to Rpb4 (and possibly other Pol II subunits) and accompanies it until imprinting. This interaction influences Pol II configuration (Figure 7A–D) and increases the likelihood of Pol II to undergo backtracking (Figure 6B). Lower panel represents non-CRAC + genes that also interact with Sfp1 (Figure 5A CONTROL, Figure 5B). We propose that promoters of non-CRAC +genes recruit Sfp1 poorly (relative to CRAC + promoters), except for small group of promoters, e.g., of RiBi genes lacking Rap1 binding site (RapBS). The dashed arrow represents this minor group. For the majority of these genes, the nuclear Sfp1 interacts directly with their elongating Pol II, as its interaction correlates with the extent of chromatin-bound Pol II (Figure 5B). This weak interaction also changes Pol II configuration (Figure 7A, ‘non-CRAC+’) and increases the propensity of Pol II to backtrack (Figure 7B). However, it either does not result in imprinting or results in rare imprinting events that went undetected by our cross-linking and analysis of cDNA (CRAC) assay. (B) Sfp1 mediates cross-talk between mRNA synthesis and decay via imprinting. The backtracked configuration, induced by Sfp1 (A), is compatible with a movement of Sfp1 from Pol II to its transcripts (see text), which is enhanced in case the GCTGCT motif is localized near Sfp1 (Figure 5C). Following co-transcriptional RNA binding, Sfp1 accompanies the mRNA to the cytoplasm and stabilizes the mRNAs. Following mRNA degradation, Sfp1 is imported back into the nucleus to initiate a new cycle. The model proposes that the specificity of Sfp1-RNA interaction is determined, in part, by the promoter (Figure 3A–B). Nevertheless, promoter binding is necessary, but not sufficient for RNA binding. See text for more details.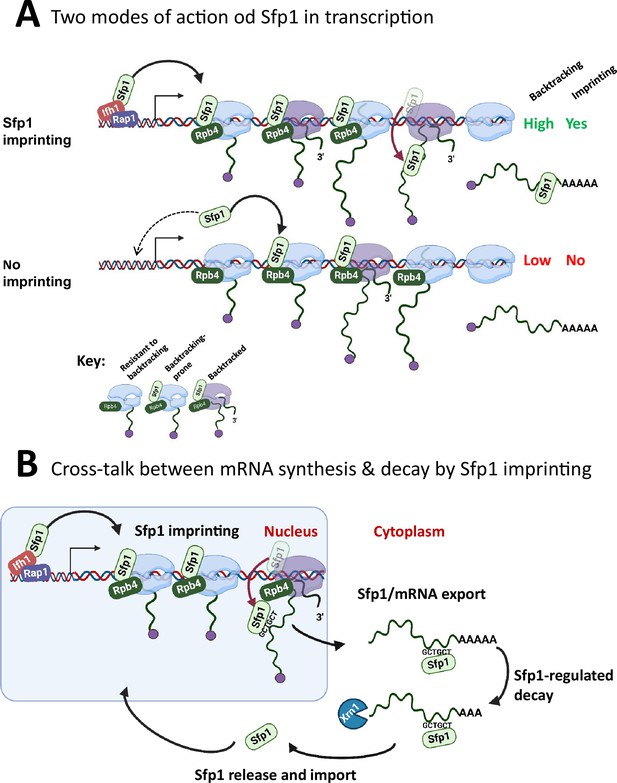
A model for split-finger protein 1 (Sfp1) functions in yeast.
The effects of Sfp1 on backtracking occur in the context of Rpb4 (Figure 7B). We note that backtracking is influenced by the alteration of Rpb4 within the elongation complex, independently of Sfp1, as the correlation between Rpb4 ChIP/Rpb3 ChIP ratios and BI is still present, albeit more mildly, in sfp1∆. Rpb4/7 was previously implicated in backtracking either by itself (Fischer et al., 2020) or in the context of the Ccr4-NOT complex (Babbarwal et al., 2014; Kruk et al., 2011). It is possible that the effect of Sfp1 on backtracking is through changing Rpb4 stoichiometry within Pol II or Rpb4 conformation. Backtracking is a conserved process across evolution that influences Pol II processivity and transcription rate (Bar-Nahum et al., 2005). We propose an additional potential function: the backtracking-prone configuration of Pol II might be more compatible with mRNA imprinting than the ‘regular’ configuration (Figure 8A). This hypothesis is inspired by the following observations: (I) Sfp1 shows a positive general influence on mRNA stability that is maximal in those mRNAs that it physically binds (Figure 4A ‘HL,’ 4B, and 6 C). (II) Sfp1 physical interaction with mRNAs occurs cotranscriptional (Figure 5C and Figure 2—figure supplement 1D). (III) The stronger the effect of Sfp1 on Pol II backtracking the stronger its effect on mRNA HL (Figure 6D). (IV) Backtracking is substantially more frequent in those genes whose mRNAs are bound by Sfp1 compared to the genome average, and significantly more dependent on this factor (Figure 6B and C). Taking all these observations together, it appears that backtracking is associated with co-transcriptional binding of Sfp1 to nascent RNA. Reassuringly, RiBi genes that exhibit Sfp1-dependent backtracking also imprint Sfp1, whereas those that do not also do not imprint (Figure 6B, ‘RiBi CRAC+,’ and ‘RiBi non-CRAC+’).
Unexpectedly, in addition to its chromatin binding feature, we find that Sfp1 binds a subpopulation of mRNAs whose transcription is stimulated by Sfp1 – ‘CRAC +mRNAs.’ CRAC + mRNAs exhibit distinctive characteristics compared to other Sfp1-regulated genes (whose mRNAs do not bind Sfp1), or other non-Sfp1-regulated genes (Figures 2A-C—7A-B, Figure 2—figure supplement 1D, F, Figure 4—figure supplement 2A-B, Figure 7—figure supplement 1A-B). Collectively, these findings demonstrate that CRAC + genes and their transcripts constitute a unique group, distinguishable from others based on several criteria, only one of them is the ability of their transcripts to bind Sfp1. CRAC + genes display a broad range of transcription rates (Figure 6A) and behave differently from the most highly transcribed genes (Figure 7A). Furthermore, while all RiBi mRNAs exhibit similar expression levels, only a subset of them binds to Sfp1. These observations collectively suggest that the 262 CRAC + constitute a distinct set of genes that unveils a new paradigm: genes controlled by a common factor both transcriptionally and post-transcriptionally, via mRNA imprinting.
We observed that RapBS-mediated promoter binding is critical for Sfp1 capacity to bind RNA (Figure 3). In RiBi genes, we found highly significant correspondence between the propensity of RiBi promoters to carry RapBS and binding of RiBi mRNAs to Sfp1 (Figure 3C). We propose that RNA-binding occurs co-transcriptionally for the following reasons: (I) Sfp1 export from the nucleus to the cytoplasm is dependent on transcription (Figure 1A), suggesting that it is exported together with the Pol II transcripts. (II) Splicing occurs co-transcriptionally (e.g. Churchman and Weissman, 2011). Consistent with co-transcriptional binding, Sfp1 binds intron-containing RPL30 RNA (Figure 2D; note that the ratio between intron-containing RNA and mature RNA is higher in the IPed lane than in the input one). (III) Binding of Sfp1 to mRNA is dependent on RapBS, suggesting that the same Sfp1 recruited to the promoter by Rap1 binds the transcript. (IV) The ChIP-exo signal drops past the GCTGCT motif in C1-C2 genes (for clusters’ definition see Figure 2B), a position where Sfp1 prefers to bind the motif containing CRAC + transcripts (Figure 5C). These observations suggest that, at least in some of their mRNA targets, Sfp1 is released from Rpb4-containing Pol II to the nascent transcripts co-transcriptionally as GCTGCT motif emerges from Pol II. However, many CRAC + do not contain detectable GCTGCT motifs or have it in a different position (Figure 2BC3 cluster). Our observation that RapBS is required to promote binding to the RPL30 mRNA sequence, a gene that contains the motif, (Figure 3B) demonstrates that motif 1 is not sufficient to recruit Sfp1. Perhaps this motif is used to stabilize the interaction. In the absence of the motif, the movement from Rpb4-containing Pol II to the emerging transcript might be related to the process of polyadenylation (see Figure 2A). Collectively, our results unveil a role for CRAC + promoters as mediators between RPB and its interacting RNAs. Whether Rpb4 dissociates together with these proteins remains to be determined. Whether the probability of mRNA imprinting is affected by the time Pol II is engaged in backtracking, until its resolution by TFIIS, is another appealing hypothesis that remains to be examined.
Binding of Sfp1 to mRNAs regulates deadenylation-mediated mRNA decay, mainly by slowing down these processes (Figure 4). Following mRNA decay, Sfp1 is imported back to the nucleus (Figure 1—figure supplement 1B, C). Following the import, Sfp1 binds to specific promoters and regulates transcription, closing the circle of gene expression regulation (Figure 8B). In this way, the synthesis and decay of Sfp1-regulated mRNAs are coordinated in a manner that maintains proper mRNA levels of a specific subset of genes.
The capacity of Sfp1 to bind mature mRNA adds additional complexity to the expression of CRAC + genes. Here, we report that Sfp1 regulates the levels of these gene products by two mechanisms: by stimulation of synthesis and by repression of decay. This raises a possible new mode of regulating mRNA level that targets the Sfp1 imprinting machinery, the extent of which would determine the mRNA HL and consequently the mRNA level. Since Sfp1 enhances both mRNA synthesis and stability, it can serve as a signaling pathway target to rapidly regulate the expression of its clients. Indeed, the expression of Sfp1 targets is highly responsive to the environment (Albert et al., 2019; Jorgensen et al., 2004).
Sfp1 was viewed as a classical TF that specifically modulates the transcription of a subset of a few hundred genes (see Introduction). We were, therefore, surprised to discover a genome-wide effects of Sfp1 depletion or deletion (Figures 4B, 6A, D, 7B–C). These results, in combination with the weak interaction of Sfp1 with chromatin (Figure 5A ‘Sfp1 Control’), are in accord with previous observation that expression of >30% of the genes, most are not considered to be direct targets of Sfp1, is affected by either Sfp1 depletion or its overexpression (Albert et al., 2019, and our unpublished NET-seq data). An important support for this notion is the correlation we found between the binding of Sfp1 to gene bodies and Pol II occupancy in all genes (Figure 5B). The general capacity of Sfp1 to alter elongating Pol II in the context of Rpb4 and to affect backtracking (Figure 7B, Figure 7—figure supplement 1A) likely underlies its widespread effect on transcription.
As mentioned above, this widespread effect of Sfp1 on backtracking correlates with higher mRNA HL in many genes (not only in CRAC+; Figure 6D). This suggests that the effect of Sfp1-dependent backtracking on mRNA HL does not always require Sfp1 binding to mRNA. Perhaps, this indirect effect of Sfp1 on mRNA HL is mediated by other factors, like Ccr4-Not, whose imprinting may also be affected by Pol II backtracking (Kruk et al., 2011; Villanyi et al., 2014; Begley et al., 2019). In contrast, the correlation between Rpb4/Rpb3 and mRNA HL is much higher in CRAC + than in the rest of the genome, and is entirely dependent on Sfp1 (Figure 7C, Figure 7—figure supplement 1B). Interestingly, we previously demonstrated that transcription elongation of Rap1-controlled genes is exceptionally affected by backtracking (Pelechano et al., 2009). We propose that Rap1-dependent recruitment of Sfp1 modulates its interaction with Pol II in a manner that permits Sfp1 transfer to mRNA (Figure 8).
Binding of Sfp1 to CRAC + mRNAs, which stabilizes the bound mRNAs, is dependent on Rap1 binding sites. On the other hand, we previously found that depletion of Rap1 leads to mRNA stabilization (Bregman et al., 2011). If the effect of Rap1 on mRNA stability is mediated only through Sfp1, we would expect that its depletion should result in mRNA destabilization, contrary to our previous observation (Bregman et al., 2011). Therefore, we conclude that Rap1 mediates additional activity that enhances mRNA decay and overrides the destabilizing effect of Sfp1, which remains to be discovered.
The Rpb4 stoichiometry within yeast Pol II is <1 (Choder and Young, 1993). It was not clear whether the sub-stoichiometric feature of Rpb4 is constant across all Pol II molecules. Here, we show that stoichiometry, as reflected by the Rpb4 ChIP/Rpb3 ChIP ratio, decreases gradually during transcription elongation. It is possible that there is a link between the Rpb4 capacity to modulate backtracking and the gradual decrease in its stoichiometry. Interestingly Sfp1 affects this stoichiometry drop, thus serving as a regulator of Rpb4 stoichiometry. Moreover, the effect of Sfp1 on mRNA stability is entirely linked to its action on Rpb4 stoichiometry (Figure 7C and Figure 7—figure supplement 1B). We note, however, that, although changes in the stoichiometry are the most likely interpretation of the change in Rpb4 ChIP/Rpb3 ChIP ratio, this change can merely reflect changes in Pol II configuration that compromise the capacity of Rpb4 to produce ChIP signal.
Interestingly, the Sfp1-mediated mechanism contrasts with that of mRNA buffering. The known model of mRNA buffering posits that a modulation of one process (e.g. transcription) is balanced by a reciprocal modulation of the other process (e.g. mRNA decay), thus maintaining the mRNA level constant, or nearly constant (Bryll and Peterson, 2023; Haimovich et al., 2013b; Pérez-Ortín and Chávez, 2022; Timmers and Tora, 2018). In contrast, the functions of Sfp1 do not result in balancing, but the contrary: the two activities of Sfp1 cooperatively increase mRNA level – increase mRNA synthesis and decrease its degradation. Thus, Sfp1 is a paramount example of enhanced gene regulation by cooperation between mRNA decay and gene transcription that we previously modeled (García-Martínez et al., 2023). Moreover, we reported that mRNA buffering maintains a constant mRNA concentration regardless of the strain growth rate, except for growth-related genes (García-Martínez et al., 2016; Chattopadhyay et al., 2022; Pérez-Ortín and Chávez, 2022). The expression of the latter genes increases with growth rate. The discovered effect of Sfp1 on its target genes, most of which are growth-related, provides a plausible mechanism to explain how growth-related genes avoid buffering when their expression must be rapidly adjusted to the ever-changing environment.
In summary, we propose that the role of certain class-specific TFs, such as Sfp1, extends beyond merely controlling various stages of mRNA synthesis and processing in the nucleus; they also regulate post-transcriptional functions in the cytoplasm. We hypothesize that a single factor (or a complex of factors) has evolved the capacity to regulate both transcriptional and post-transcriptional functions to facilitate cross-talk between these two mechanisms. Sfp1 impacts the mRNA life cycle – from transcription to degradation. Interestingly, the Sfp1 effect on transcription elongation is linked to its function on mRNA stabilization, suggesting that all its functions are connected. Sfp1 possesses two classical zinc fingers. Given that proteins with zinc fingers, characteristic of many TFs, are known to be involved in DNA, RNA, and protein binding, it is conceivable that this zinc finger-containing protein is suitable for participating in the mRNA imprinting process, as proposed here (Figure 8B). Given that RBPs have been proposed to perform multiple roles in RNA-based regulation of gene expression in mammals (Xiao et al., 2019), we anticipate that the case of Sfp1 as an imprinting factor will serve an example of this important type of gene regulators in other eukaryotes.



Add Comment