A higher M1/M2 macrophage ratio indicates antitumor immunity in melanoma
TAMs play a critical role in shaping the immune status of tumors. Responding to the cues from different TME cells, TAMs exhibit a broad spectrum of phenotypes, ranging between M1 and M2 macrophages or even adopting hybrid phenotypes [15]. To understand the role of macrophages in melanoma TME, we first evaluated the composition of M1-like and M2-like macrophages in human and murine melanoma tumors. High expression of M1 markers such as CD38 and CD80 and high infiltration of M1-like macrophages were associated with improved overall survival in the TCGA SKCM dataset (Fig. 1A-B), whereas high expression of M2 markers ARG1 and CD163 and high M2-like macrophage tumor infiltration was associated with poor survival (Fig. 1C-D). We further validated the role of macrophages in C57BL/6 mice engrafted with SM1 melanoma tumors and evaluated for correlation between macrophage phenotypes and tumor growth. At the endpoint, 25 days post-tumor implantation, we assessed the infiltration of M1 (F4/80+, Cd80+) and M2 macrophages (F4/80+, Cd206+), plotted as a percentage of total live cells. Figure 1E represented the tumor growth kinetics, and Fig. 1F indicated that antitumor M1 macrophages negatively correlated with tumor volume, whereas protumor M2 macrophages positively correlated with tumor volume (Fig. 1G). Figure 1H revealed a negative correlation between the M1/M2 ratio and tumor growth, which has also been observed in clinical studies [16], suggesting that the SM1 murine melanoma model is ideal for investigating the role of macrophages in tumor progression. Overall, our data indicate that TAMs polarized towards M2 phenotype play a critical role in the progression of both human and murine melanoma tumors.
M1/M2 ratio reflects the immune status of the tumor microenvironment. (A–D) Kaplan Meier survival analysis of key M1 macrophage genes CD38, CD80, and M2 macrophage markers ARG1, CD163 with overall survival in the cancer genome atlas (TCGA) skin cutaneous melanoma patients (SKCM) (n = 470). High or low gene expression of macrophage phenotype markers is correlated to increased or decreased presence of tumor associated macrophages (E) Tumor growth chart representing the growth kinetics of SM1 murine melanoma tumors with BrafV600E mutation in immunocompetent C57BL/6 mice at the tumor volume endpoint which was 25 days post-tumor implantation (20 mm diameter). (n = 20 mice) (F) Negative correlation between SM1 murine melanoma tumor volume and tumor-associated M1 macrophages (F4/80 + Cd80 + as % of live cells). (G) Positive correlation between the tumor volume and tumor-associated M2 macrophages (F4/80 + Cd206 + as % of live cells). (H) Negative correlation between the tumor volume and M1/M2 macrophage ratio
HDAC6 inhibition affects macrophage phenotype and function both in vitro and in vivo
Previous studies proved that HDAC6i treatment reduced tumor growth and is associated with an enhanced M1/M2 ratio [11, 12, 14]. In the SM1 murine melanoma model, intraperitoneal administration of the HDAC6i NexturastatA (NextA) resulted in reduced tumor size compared to the vehicle-treated group (Fig. 2A). Phenotyping of TAMs by flow cytometry showed a slight increase in M1 macrophages and a significant decrease in M2 macrophages, thus effectively increasing the M1/M2 ratio in the NextA treated group (Fig. 2B). Similarly, BMDMs treated with NextA indicated an increase in M1 polarization and significant reduction of M2 polarization, consistent with in-vivo observations (Fig. 2C).
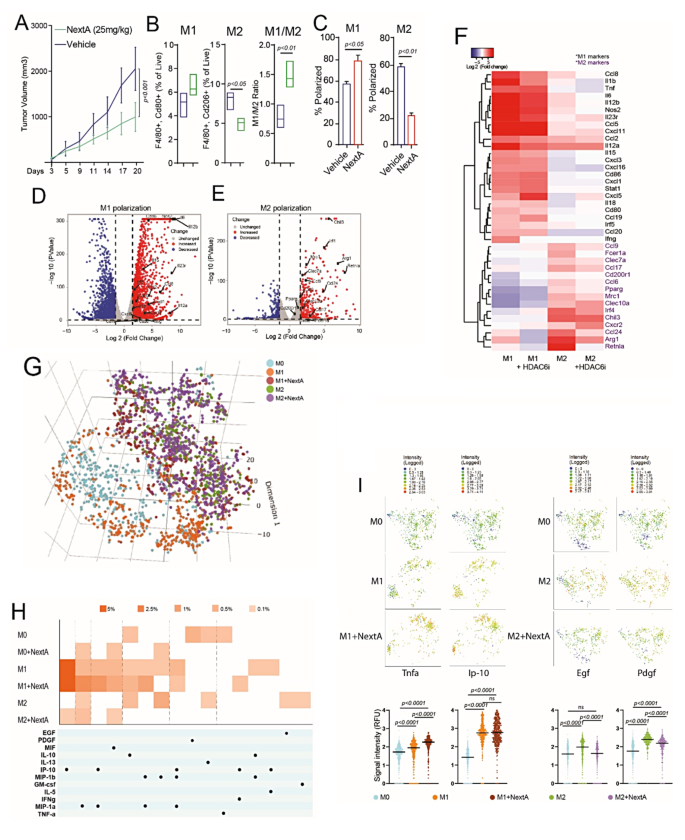
HDAC6 inhibition regulated M2 macrophage phenotype. (A) Tumor growth chart of syngeneic SM1 murine melanoma tumors in C57BL/6 mice treated with intraperitoneal administration of 25 mg/kg HDAC6 inhibitor, NexturastatA (NextA) or vehicle. (B) Tumor-associated M1 (F480 + Cd80+) and M2 (F4/80 + Cd206+) macrophages as a % of live cells, respectively, and M1/M2 ratio in vehicle or NextA treated mice bearing SM1 murine melanoma tumors. Tumors were collected on day 20 with the tumor size endpoint as 20 mm diameter. (C) Polarization efficiency of murine bone marrow-derived macrophages (BMDMs) to M1 and M2 phenotypes after treatment with HDAC6i, NextA (5µM) determined by flow cytometry. (D) Volcano plot showing fold-change and p-value for comparing vehicle-treated M1 versus M0. M0 are naïve macrophages derived from the mouse bone marrow. (E) Volcano plot showing fold-change and p-value for the comparisons and vehicle-treated M2 versus M0 macrophages. The significance level was determined by log2 fold changes ≥ 1.5 (upregulation/increased) or ≤-1.5 (downregulation/decreased) and p-value < 0.05. Differentially expressed genes are depicted in blue and red, where M1 and M2 markers are labeled in figures (D) and (E), respectively. (F) Heatmap of known markers for classically activated M1 and M2-like macrophages. Differential expression for HDAC6 inhibition versus vehicle was performed, and M1 markers (black) and M2 markers (purple) were represented using the log 2 transformed fold changes relative to vehicle-treated M0. (G) We analyzed about 659 M0, 407 M1, and 450 M2 macrophages treated with vehicle; 424 M1 and 378 M2 macrophages treated with NextA on mouse innate immune IsoCode chips on the Isoplexis platform. Uniform manifold approximation and projection (UMAP) dimensionality reduction analysis of BMDMs at a single cell resolution separated them into defined clusters based on their secretome profile. (H) A polyfunctionality heatmap representing each macrophage phenotype secretes more than one cytokine/chemokine. The intensity of orange squares in the heatmap represents the percentage of cells secreting the cytokine/chemokine indicated by corresponding black dots below. (I) 2-D tsne plots where each cell is represented as a colored dot. As shown in the intensity scale, blue indicates low expression, and red indicates high expression. Bar graphs of respective cytokines are represented as signal intensity. Proinflammatory cytokines Tnfa, and T-cell recruiting chemokine Ip-10 (Cxcl10) are elevated in M1 + NextA, whereas growth factors Egf and Pdgf secreted by M2 phenotype are decreased with NextA treatment
Transcriptional regulation of macrophage phenotypic markers by HDAC6i has not yet been reported, so we performed bulk RNA-seq analysis from BMDMs, including M0, M1, and M2 macrophages treated with vehicle or NextA. Volcano plot of gene expression changes in Fig. 2D represented an increased expression of M1 markers and proinflammatory genes, including Nos2, Cd86, Il12a, Il12b, Ifng, Cxcl1, Cxcl3, and Il6. On the other hand, M2 polarization upregulated M2 markers and anti-inflammatory genes in Fig. 2E, including Arg1, Retnla, Mrc1, Chil3, Ccl17, Ccl24, and Irf4. Thus, transcriptomic analysis confirmed that gene expression profiles of polarized macrophages correlated with previously reported studies [17]. The bar plot shows an exclusive expression of M1-specific genes (Supp. Fig. S1A) and M2-specific genes (Supp. Fig. S1B) in respective phenotypes, represented as heatmaps in Supp. Fig. S1C. Unsupervised cluster analysis of phenotypic markers between vehicle and NextA-treated M1 and M2 macrophages indicated two major clusters in the heatmap, as shown in Fig. 2F. One cluster included proinflammatory M1 genes wherein expression was maintained or slightly increased with NextA. In the other cluster, expression of M2 markers such as Retnla, Arg1, and Mrc1 was decreased with NextA, suggesting a global suppression of M2-associated gene signature. Further, pathway analysis of differentially expressed genes indicated a significant downregulation of anti-inflammatory cytokine IL10 signaling with NextA treatment of M1 macrophages (Suppl. Fig. S1D), potentially enhancing the proinflammatory nature of M1 macrophages. In M2 macrophages, NextA treatment suppressed several cell cycle and cell proliferation-related pathways (Suppl. Fig. S1F).
To validate the functional consequence of HDAC6 inhibition on macrophage function, we performed a single-cell secretome (sc-secretome) analysis of BMDMs treated with vehicle or NextA. UMAP analysis of single cells in Fig. 2G separated macrophages into distinct clusters based on their secretome profiles. Figure 2H is a polyfunctional heatmap of macrophages capable of secreting more than one cytokine or chemokine, indicating the percentage of M1 and M1 + NextA macrophages secreting chemokine Ip-10 (Cxcl10), Mip-1a (Ccl3), Mip-1b (Ccl4), and Mif. In M2 macrophages, NextA reduced the percentage of macrophages secreting growth factor Egf and anti-inflammatory cytokines Il-10 and Il-13 while increasing the percentages of M2 macrophages secreting Ip-10, Mip-1a, and Mif, suggesting a shift away from M2 function. 2D-tsne plots indicated clusters of cells with increased expression of inflammatory cytokines Tnfa and T-cell recruiting Ip-10/Cxcl10 in M1 + NextA compared to M1 or M0 macrophages (Fig. 2I). Conversely, NextA treatment in M2 macrophages suppressed M2 function by decreasing the secretion of growth factors such as Egf and Pdgf compared to M2 macrophages. Thus far, the data from in-vivo study, transcriptomic analysis, and sc-secretome analysis of BMDMs comprehensively demonstrated that HDAC6 inhibition profoundly affected the macrophage phenotype and function.
HDAC6 inhibition predominantly affects M2 macrophages
TAMs predominantly exhibit M2 phenotype [18]. Therefore, suppressing M2 macrophages could benefit cancer patients. Towards this approach, we investigated how HDAC6i could modulate M2 macrophage phenotype and function. Gene expression analysis of M2 markers indicated a significant decrease in the expression of Arg1, Mrc1, and Tgf1 (Fig. 3A) in NextA-treated BMDMs, further validating the transcriptomics data. Of note, NextA did not induce cytotoxicity on murine BMDMs even at concentrations as high as 10µM (Supp. Fig. S2A). Therefore, the suppression of tumor growth in vivo was predominantly due to the immunomodulatory effects of NextA on the TME. Importantly, other HDAC6is have been shown to have minimal cytotoxic effects in normal and transformed cells [14].
Next, Thp1-derived human macrophages were treated with vehicle or NextA before being polarized to M1 or M2. As shown in Fig. 3B by qRT-PCR analysis, NextA significantly decreased the expression of M2 markers, MRC1 (CD206), and CD209 but minimally affected the polarization of M1 macrophages (CD80 and CD86). A similar reduction of CD206 expression was observed with other HDAC6is tubacin and tubastatin A (Supp. Fig. S2B). In addition to pharmacological inhibition, we used a genetic approach with shRNA to knockdown Hdac6 in murine BMA3.1A7 macrophages. Immunoblot analysis indicated partial knockdown of Hdac6 protein in BMA3.1A7 cells (Fig. 3C), as shown by increased acetyl-tubulin, a known substrate to be deacetylated by HDAC6 [19]. Gene expression analysis of the M2 markers, Arg1, Mrc1, and Tgf1 by qRT-PCR revealed a reduction in the expression in Hdac6 knockdown (HDAC6KD) compared to non-target macrophages (Fig. 3D). Flow cytometry analysis indicated a decrease in M2 polarization which was comparable to NextA treatment (Fig. 3E). Evaluation of NextA-treated macrophages by immunoblot analysis of polarization markers, indicated a decrease in arginase 1 (Arg1) in M2 macrophages and no effect on induced nitric oxide synthase (iNOS) in M1 macrophages (Fig. 3F). Furthermore, analysis of M2 macrophages by immunofluorescence demonstrated a significant decrease in the M2 marker, Cd206 compared to vehicle treatment (Fig. 3G). Taken together, pharmacological or genetic inhibition of HDAC6 in both human and murine macrophages significantly decreased M2 polarization, demonstrating that HDAC6 inhibition has a substantial immunomodulatory effect on macrophages.
HDAC6 inhibition suppressed macrophage polarization towards the M2 phenotype. (A) Real time quantitative PCR (qRT-PCR) analysis of mRNA expression levels of M2 markers Arg1, Tgfb1, and Mrc1 in vehicle and NextA (5µM) treated murine BMDMs. (B) Analysis of mRNA expression levels of M2 markers MRC1 (CD206) and CD209, M1 markers CD80 and CD86 in vehicle and NextA (5µM) treated human macrophages derived from Thp1 monocytic cell line. (C) Immunoblot analysis of Hdac6 after non-target or Hdac6 targeted shRNA mediated knockdown in murine macrophage cell line BMA3.1A7. Tubulin is protein loading control. (D) Analysis of mRNA expression levels of M2 markers Arg1, Mrc1, and Tgf1 by qRT-PCR in non-target and Hdac6 knockdown (HDAC6KD) BMA3.1A7 cells by q-PCR. (E) Mean fluorescence intensity (MFI) of CD206 in HDAC6 knock down (HDAC6KD) BMA3.1A7 murine M2 macrophages analyzed by flow cytometry. (F) Immunoblot analysis of BMDM macrophages for M1 (iNOS) and M2 (Arg1) markers. Tubulin is a loading control, and acetyl-tubulin is a marker for HDAC6 inhibition. (G) Immunofluorescence analysis of M2 marker CD206 in naïve (M0) and M2 polarized BMDMs with or without NextA treatment (5µM). (H) Immunoblot analysis of IL6 mediated STAT3 phosphorylation at tyrosine 705 (Y705) in RAW264.7 murine macrophages treated with NextA (5µM) or vehicle. (I) Flow cytometry analysis of BMDM derived M2 macrophages as a percentage of Cd45 + cells treated with HDAC6 inhibitor, NextA (5µM) and STAT3 inhibitor, Stattic (10µM). (J) Analysis of mRNA expression of Arg1, Tfgb1, Retnla (Fizz1) by qRT-PCR in bone marrow-derived M2 macrophages treated with NextA (5µM) and Stattic (10µM). (K) Violin plot representing the expression of STAT3 target genes in M1 and M2 macrophages treated with NextA shown as fold change relative to M0. Data obtained from transcriptomics analysis of murine BMDMs. (L) Macrophage repolarization assay, mRNA expression analysis of Arg1 by qRT-PCR in M1 macrophages exposed to M2 polarizing cytokines
We and other groups previously established that HDAC6 interacts with STAT3 to regulate the expression of STAT3 target genes [20,21,22]. Moreover, STAT3 is reported to regulate the expression of Arg1 in myeloid-derived suppressor cells [23]. Therefore, we reasoned that STAT3, a key transcription factor promoting the M2-like phenotype, might regulate Arg1 expression in macrophages. Immunoblot analysis of RAW264.7 macrophages treated with IL-6 cytokine indicated STAT3 phosphorylation at tyrosine-705 (Y705), and NextA treatment decreased STAT3-Y705 phosphorylation, which aligned with our previous report [20] (Fig. 3H). Flow cytometry analysis of M2-polarized BMDMs treated with NextA or the STAT3 inhibitor, Stattic, showed a significant decrease in M2 macrophages (Fig. 3I). The expression of M2 markers, including Arg1, Tgf1, and Retnla (Fizz1) in BMDMs was significantly decreased in M2 macrophages treated with NextA (5µM) and Stattic (10µM) (Fig. 3J). Interestingly, NextA significantly affected Arg1 and Retnla (Fizz1) gene expression more than STAT3 inhibition. As shown in Suppl. Fig. S2C, the expression of immunosuppressive cytokine Il10 upregulated in M1 macrophages as negative feedback to the inflammatory response mediated by IFNγ and LPS was decreased with NextA treatment. Furthermore, Socs3 expression, a negative regulator of STAT3 signaling, was upregulated in the NextA-treated M1 macrophages, which explains the downregulation of Il10 expression. Further analysis of STAT3 target genes in M1 and M2 macrophages from RNA-seq analysis indicated that STAT3 targets in M2 macrophages were significantly suppressed than STAT3 targets in M1 macrophages (Fig. 3K). ). In addition, gene set enrichment (GSEA) analysis of M2 and M2 + NextA transcriptomes indicated a significant number of genes in the IL6-JAK-STAT3 pathway were downregulated (Suppl. Fig. S3A). STAT3 target genes in M1 and M2 macrophages treated with NextA are shown in Suppl. Table. S5. For the first time, we report that HDAC6 inhibition can affect M2 macrophage polarization through STAT3 pathway suppression.
We further performed a repolarization assay where M1-like macrophages were treated with vehicle or NextA and subsequently exposed to M2-polarizing cytokines Il-4 and Il-13. M2 polarization was demonstrated by increased M2 marker expression, Arg1 by qRT-PCR (Fig. 3L, compare M0 vs. M2). Consistently, NextA diminished M2 polarization (Fig. 3L, compare M0 vs. M2 + NextA). Upon repolarization with M2 cytokines, Arg1 expression significantly increased in vehicle-treated M1 macrophages but not in the NextA-treated M1 macrophages (Fig. 3L, compare M0 vs. M1→M2 vs. M1 + NextA→M2), suggesting that M1 macrophages treated with NextA exhibit a lock into the M1 phenotype. A similar effect was demonstrated using RAW 264.7 macrophages where expression of the M2 marker, Mrc1 was suppressed in M1 + NextA macrophages upon repolarization with M2 cytokines compared to M1-like macrophages (Suppl. Fig. S2D). Taken together, either pharmacological or genetic inhibition of HDAC6 in human and murine macrophages decreased polarization of macrophages towards tumor-promoting M2 phenotype while retaining the M1 phenotype, suggesting that HDAC6is could be used as therapeutic immunomodulatory agents to control the macrophage phenotype and function.
HDAC6 inhibitor treated macrophage-based adoptive cell therapy improved antitumor immune response in melanoma
HDAC6 inhibition in macrophages decreased the M2 phenotype while retaining the M1 phenotype, offering an ideal adoptive cell therapy strategy (ACT). Therefore, we tested this approach with BMDMs treated ex-vivo with NextA and then transplanted into SM1 tumors. ACT is shown in the schematic in Suppl. Fig. S2E where macrophage ACT only resulted in a significant tumor growth reduction with NextA-treated M1 macrophages but not with vehicle-treated M1 macrophages. M0 and M2 macrophage transplantation with NextA did not reduce the tumor size significantly. Encouraged by the tumor reduction with HDAC6-inhibited M1 macrophages, we expanded our macrophage ACT to a larger cohort of mice at time points shown in the schematic in Fig. 4A. Intratumor dose titration studies indicated that 5 mg/kg of NextA (Suppl. Fig. S2F), and 1 × 106 M1 + NextA treated macrophages (data not shown) were optimal to elicit a tumor reduction response. Furthermore, a cytotoxicity assay performed with NextA indicated minimal toxicity at 5µM compared to Panobinostat (LBH-589), which is a pan-HDAC inhibitor inducing significant cytotoxicity at comparable NextA concentration (Suppl. Fig. S2G). Therefore, any potential tumor reduction from NextA intratumor administration is due to tumor immune modulation rather than a direct cytotoxic effect. Consistent with Suppl. Fig. S2E, intra-tumor ACT of NextA-treated M1 macrophages resulted in a significant tumor size reduction compared to other treatment groups (Fig. 4B). NextA treatment performed better than vehicle-treated M1 macrophage ACT and the control group. The tumor reduction also translated into better survival with M1 + NextA followed by NextA and M1 groups, as shown by the Kaplan-Meier survival plot in Fig. 4C. More importantly, we also observed that ACT with BMDMs derived from HDAC6 knockout mouse recapitulated the results of M1 + NextA (Fig. 4D). Overall, ACT with NextA-treated M1 macrophages significantly reduced tumor volume and improved survival compared to the control group, highlighting the efficacy of macrophage ACT.
Adoptive cell therapy of HDAC6 inhibitor treated M1 macrophages diminishes melanoma tumor growth in immunocompetent mice. (A) Schematic workflow of the macrophage adoptive cell therapy (ACT) in C57BL/6 syngeneic SM1 murine melanoma model. (n = 10 mice/group) (B) Tumor growth chart of SM1 melanoma tumors treated with vehicle (PBS), intratumor ACT of M1 (1 × 106) macrophages, injection of NextA (100ug), and intratumor ACT of M1 macrophages (1 × 106) pretreated with NextA (5µM) ex-vivo. (C) Survival analysis of mice treated with macrophage ACT. (D) Tumor growth chart of SM1 murine melanoma tumors with vehicle (PBS) or bone marrow-derived M1 macrophages from HDAC6KO mouse. (E) Flow cytometry-based immunophenotyping of M1, M2 macrophages as a fraction of F480 + Cd80 + and F4/80 + Cd206 + macrophages, respectively, and M1/M2 ratio in SM1 murine melanoma tumors treated with macrophage ACT. (F) CD8 T-cells as a fraction of Cd3 + cells. (G) CD4 T-cells, and T-regs as a fraction of Cd3 + cells. (H) Analysis of SM1 murine melanoma tumors treated with vehicle (Control), M1 macrophages, or HDAC6 inhibitor treated M1 macrophages (M1 + NextA) by hematoxylin and eosin (H&E) staining. Tumor stroma is indicated with white arrows. (I) Transplanted macrophages were derived from the bone marrow of UBC-GFP mice with ubiquitous GFP expression, enabling us to detect them by immunostaining with an anti-GFP antibody. (J–K) Immunohistochemistry staining to detect GFP-expressing macrophages, GFP and Cd38 expressing macrophages, GFP and Cd206 expressing M2 macrophages. White arrowheads indicate transplanted GFP macrophages. GFP expression is shown in black color. Cd38 and Cd206 expression are represented in brown color
Flow cytometry analysis indicated a sharp increase in M1 macrophages and no change in M2 macrophages in the M1 + NextA ACT group compared to Vehicle and M1 macrophage ACT treatment groups. This increase in the M1 macrophages from ACT translated into a higher M1/M2 ratio in the M1 + NextA group. (Fig. 4E). The Higher M1/M2 ratio was further associated with an increased infiltration of CD8 + effector T-cells in the M1 + NextA group (Fig. 4F). There was also an increase in effector memory (EM) cells and central memory (CM) CD8 T-cells. Despite a trend towards an increase in CD4 + cells, there was no significant difference in the immunosuppressive regulatory T-cells (T-regs) among the treatment groups (Fig. 4G). Overall, an increase in the number of pro-inflammatory immune cells, including M1 macrophages and CD8 effector T-cells, suggested that macrophage ACT activated both innate and adaptive antitumor immunity.
Histological analysis of tumor sections from vehicle, M1, and M1 + NextA treatment groups (Fig. 4H) showed areas of increased stroma (white arrows) in the M1 and M1 + NextA tumor sections compared to the vehicle-treated group, suggesting there may be an overall increase in the infiltration of immune cells. Notably, GFP-positive M1 and M1 + NextA macrophages (stained black) were still viable two weeks post-transplantation (Fig. 4I). Further analysis of M1 marker Cd38 (stained brown) visibly indicated an increased presence of M1 macrophages in the proximity of GFP macrophages in both M1 and M1 + NextA groups. Importantly, some GFP macrophages stained for both GFP and Cd38 (white arrowheads), demonstrating that transplanted M1 + NextA macrophages retained the M1 phenotype (Fig. 4J). On the other hand, M2 tumor macrophages were reduced in the vicinity of M1 GFP macrophages (as indicated by white arrowheads) treated with NextA, suggesting a proinflammatory TME in the M1 + NextA group (Fig. 4K). Overall, HDAC6 inhibition in ACT macrophages significantly reduced tumor volume through antitumor M1 macrophages and increased infiltration of CD8 effector T-cells.
Single-cell profiling of tumor immune infiltrate reveals a proinflammatory TME post macrophage ACT
scRNA-seq analysis of myeloid cells across multiple cancers revealed the diversity of TAMs ranging from inflammatory-proliferating to regulatory TAMs [4]. Therefore, we explored the cellular composition of SM1 tumors post-ACT using the 10X Genomics platform. UMAP analysis [24] of sorted Cd45 + cells isolated from tumors indicated distinctive clusters of immune cells from Control and M1 + NextA tumors. SingleR analysis [25] revealed major clusters including macrophages, monocytes, and T-cells (Fig. 5A). Further analysis of macrophages resolved into four distinct subclusters Mac1, Mac2, Mac3, and Mac4 (Fig. 5B). The control tumor was predominant with Mac2 and Mac4 subclusters; whereas the M1 + NextA tumor group was enriched with Mac1 subcluster. Gene expression analysis indicated the Mac1 subcluster expressed an inflammatory gene signature and a Mac2 subcluster expressed an anti-inflammatory signature akin to M1 and M2 macrophages, respectively. Therefore, ACT with M1 + NextA macrophages increased the percentage of M1-like macrophages with a concomitant decrease of M2-like macrophages (Fig. 5B insert). Feature plots of M1-like genes Cxcl9, Cd72 (Fig. 5C), CD80, and CD86 (Suppl Fig. S4A) indicate expression in the Mac1 subcluster of M1 + NextA tumors. Conversely, M2 genes, Arg1 and Mrc1 (Fig. 5D) and Thbs1 (Suppl Fig. S4B) were mostly expressed by the Mac2 subcluster in the control tumors. Differential gene expression analysis of macrophage subclusters enriched for Aif1, Cxcl9, Cd72, Lst1, Hck, and Pou2f2 gene signature in the Mac1 that are linked to macrophage activation, phagocytosis, T-cell recruitment and inflammatory response [26], [27]. Overall, the Mac1 subcluster was similar to the inflammatory TAMs reported by Ma et al. [4]. Genes enriched in the Mac2 subcluster included Bnip3, Egln3, Hilpda, Slc2a1, Tpi1, Aldoa, Tmem189, Fnip2, and Anxa2 and its gene signature was similar to the regulatory TAMs reported by Ma et al. [4]. We observed that the Mac1 gene signature was partially expressed by the Mac4 subcluster, while the Mac2 gene signature was partially expressed by the Mac3 subcluster, as shown in the bubble plot in Fig. 5E and the heatmap in Fig. 5F. We further interrogated the expression of M1 and M2 gene signatures identified from bulk RNA-seq (Fig. 2F) and found that M1 gene signature was upregulated in the ACT tumor macrophages (Suppl Fig. S4C) and M2 gene signature was upregulated in the Control tumor macrophages (Suppl. Fig. S4D). The data thus far indicated that M1 + NextA macrophage ACT resulted in an increased M1/M2 ratio, and the expression of M1-associated gene signature is positively correlated to survival benefits to patients.
Macrophage ACT activated inflammatory immune responses in the TME. (A) UMAP analysis of Cd45 + sorted tumor infiltrated immune cells from Control and M1 + NextA macrophage treated tumors analyzed with 10xGenomics workflow. UMAP cluster analysis of aggregated immune cells by SingleR cell annotation revealed major immune cell populations. (B) Tumor macrophage subclusters Mac1, Mac2, Mac3, and Mac4 show differential presence represented as a percentage of total tumor macrophages (insert with stacked bar plot). (C) Feature plots representing the expression of inflammatory genes Cxcl9 and Cd72 associated with M1 phenotype in Mac1 macrophage subcluster in Control and M1 + NextA tumors. (D) Feature plots representing expression of tumor promoting genes Arg1 and Mrc1 associated with M2 phenotype in Mac2 macrophage subcluster in Control and M1 + NextA tumors. (E) Bubble plot showing differentially expressed top significant gene between macrophage subcluster. Mac1 associated genes are shown in orange, and Mac2 associated genes are shown in green color. (F) Heatmap representing the expression of Mac1 and Mac2 gene signatures in single cells across 4 macrophage subclusters. (G) Circos plot showing potential interactions between ligands expressed by macrophages in Mac1 and Mac2 subclusters and receptors expressed on the receiver T-cells. Heatmap representing the pathway analysis of activated target genes in receiver monocytes. MSigDB Hallmark 2020 module in Enrichr was used to generate the heatmap. (H) Circos plot showing potential interactions between ligands expressed by macrophages in Mac1 and Mac2 subclusters and receptors expressed on the receiver monocytes and corresponding heatmap of pathway analysis
To further understand the impact of NextA-treated M1 macrophage ACT on the TME, we performed sc-secretome profiling of TAMs on the Isoplexis platform using a mouse innate immune IsoCode chips. The 3D-tsne plot indicated the clustering of TAMs from control tumors separated from ACT-treated tumors based on their secretome profiles (Fig. 6A). Compared to BMDMs, TAMs were relatively more polyfunctional, as indicated by the number of secreted cytokines and chemokines shown in heatmap, potentially due to a niche of cell-cell communication that is absent in BMDMs cultured in-vitro (Fig. 6B). The majority of TAMs from both control and ACT tumors secreted inflammatory cytokines, as shown in the polyfunctional strength index plot in Fig. 6C. Expression of cytokines and chemokines by TAMs are shown as 2-D Tsne plots in Fig. 6D. Compared to the control tumor, TAMs from the ACT tumor had increased secretion of inflammatory factors including Tnfa, Mif, T-cell recruiting chemokine Ip-10, and Il-5, whereas decreased secretion of anti-inflammatory Il-10. Pleotropic cytokines Il-6 secretion was increased, while Mip-1b (Ccl4) was reduced with ACT TAMs (Fig. 6E). Thus, the sc-secretome data indicated that ACT with NextA-treated M1 macrophages altered the function of TAMs towards an inflammatory phenotype.
Single-cell secretome analysis indicated polyfunctionality of tumor macrophages. Two weeks post-ACT with NextA-treated M1 macrophages, tumors were harvested and flow-sorted for TAMs with Cd45 and F40/80 antibodies. About 247 TAMs from the control group and 585 TAMs from the M1 + NextA ACT group were analyzed with high IsoQ scores. (A) A 3D-tsne plot of Cd45 + F4/80 + sorted tumor macrophages from Control and M1 + NextA macrophage ACT treated tumors analyzed on IsoSpark. (B) Polyfunctionality heatmap representing the percentage of cells expressing different cytokines and chemokines by tumor macrophages from Control and M1 + NextA ACT tumors. (C) Polyfunctionality strength index of TAMs expressing cytokines and chemokines. (D–E) 2-D Tsne plots and scatter plots of signal intensities of represented secreted factors from TAMS isolated from Control and M1 + NextA ACT tumors
Ligand-receptor interaction analysis indicated immune activating signals upregulated with macrophage ACT niche
Cell-to-cell communication through secretory factors and ligand-receptor interactions sustains the dynamic nature of the TME. We previously reported that a functional immune system is required for HDAC6 inhibition-mediated tumor suppression using immunodeficient SCID mice and CD4/CD8 T-cell depletion assays [28]. This was particularly evident from sc-secretome analysis, where TAMs were relatively more polyfunctional compared to in vitro cultured BMDMs due to the interactive nature of the TME. Therefore, we interrogated whether macrophage ACT affected cell-cell interactions through differential NicheNet analysis. The top 30 ligand-receptor interactions were represented as a heatmap (Suppl. Fig. S5A) showing potential interactions between macrophage ligands and T-cell receptors. In the M1 + NextA niche, ligand-receptor interaction analysis indicated that Cd72 in the Mac1 subcluster potentially interacted with Sema4d and Cd5, expressed on T-cells in M1 + NextA tumors. Cd72 is expressed in a subset of inflammatory macrophages [29], while Sema4D (Cd100) [30] and Cd5 [31] are highly expressed in T cells. Similarly, T-cell recruiting Cxcl9 and Cxcl10 chemokines were upregulated in the Mac1 subcluster, potentially recruiting T-cells into M1 + NextA -treated tumors through interaction with Cxcr3. Increased expression of Cxcl10 (Ip-10) was validated by sc-secretome analysis of TAMs isolated from M1 + NextA tumors (Suppl. Fig. S4E). Next, we analyzed ligand activity based on the expression of target genes in receiver cells. Il27a and Ebi3 (Il27b) were the most active among the ligands, with the ability to activate most of the top target genes (Suppl. Fig. S5B). The circos plot in Fig. 5G summarizes potential interactions between the ligands and receptors expressed on the Mac1 and Mac2 macrophage subclusters and tumor-infiltrated T-cells. Pathway analysis of top target genes in T-cells indicated activation of allograft rejection and interferon-gamma response among the significant pathways based on p-values. Overall, NicheNet analysis demonstrated macrophage-mediated T-cell activation.
Since monocytes are the major contributors to the macrophage population in tumors, we interrogated the macrophage-monocyte interactions using NicheNet analysis. Similar to T-cells, the interaction of ligands Cd72, Cxcl9, and Cxcl10 from Mac1 was also observed in monocytes. Furthermore, monocyte recruiting chemokine Ccl5 from Mac1 potentially interacts with Ccr1 in Control tumor monocytes. Also, this interaction was upregulated with Ccl5 in M1 + NextA tumor monocytes (Suppl. Fig. S5C). In addition, Ccl12 from both Mac1 and Mac2 subclusters interact with Ccr2 in M1 + NextA tumor monocytes. Adam17, Ltb, Ptprc, Ly86, Ccl12, and Tnfsf13b appear to be the most active ligands potentially upregulating most top-scored target genes, including Stat1 transcription factor (Suppl. Fig. S5D). Pathways analysis of top target genes in monocytes reflected that interferon-gamma and interferon alpha response were among the significant pathways based on p-values. The circos plot in Fig. 5H summarizes potential interactions between the ligands and receptors expressed on the Mac1 and Mac2 macrophage subclusters and tumor-infiltrated monocytes.
In addition, we analyzed the fate of monocytes by trajectory analysis (Suppl. Fig. S6A). Macrophages in the control tumor clustered as several subsets transitioning from infiltrated monocytes. On the other hand, M1 + NextA tumor macrophages clustered exclusively into a dichotomous grouping. Pathway analysis of top 100 significant genes in Node 1 macrophages was enriched with oxidative phosphorylation, suggesting a more M2 phenotype (Suppl. Fig. S6B). Node 2 macrophages were enriched with inflammatory response pathways, which is evident by macrophage activation GO biological process demonstrating more M1-like characteristics (Suppl. Fig. S6C). The Node 3 was enriched with Il-2/Stat5 signaling pathway and Jak-Stat receptor signaling biological process representing an intermediate phenotype between M1 and M2 macrophages (Suppl. Fig. S6D). More importantly, in M1 + NextA tumors, transitioning monocytes appear closely aligned to Node 2 inflammatory M1-like macrophages compared to the control tumor macrophages. Overall, computation analyses indicated that Mac1 subcluster from differential NicheNet and Node2 macrophages from pseudotime analysis have a proinflammatory and activating effect on tumor-infiltrated T-cells and monocytes. This agrees with the biological responses such as increased M1/M2 ratio and increased CD8 T-cell infiltration with macrophage ACT in vivo studies.
Finally. we performed an in-vitro co-culture assay of T-cells and macrophages with or without NextA treatment (5µM) for 72 h to validate macrophage-mediated T-cell activation. Analysis of supernatant for cytokine and chemokine secretome indicated an increased secretion of proinflammatory factors such as IP-10 (Cxcl10), IFNg, TNFa, and RANTES by T-cell co-cultured with M1 + NextA macrophages (similar to M1 + NextA tumor macrophages) than T-cells cultured with M1 macrophages (Suppl. Fig. S6E). Furthermore, the proliferation of CD8 T-cells suppressed when co-cultured with M2 macrophages was rescued with NextA-treated M2 macrophages (Suppl. Fig. S6F), suggesting that HDAC6 inhibition can influence T-cell activity through macrophages.
Macrophage ACT in humanized mice reduced melanoma tumor growth
Next, we used human macrophages derived from circulating monocytes to demonstrate the translational potential of macrophage-based ACT in treating cancers. Sc-secretome analysis of human M1 or M2 macrophages treated with NextA revealed well-defined clustering of macrophages in a 3D-tsne plot, suggesting that each macrophage phenotype has a unique secretome profile (Suppl. Fig. S7A). Polyfunctional heatmap indicated that NextA treatment increased the percentage of polyfunctional M1 macrophages secreting more than one cytokine or chemokine (Suppl. Fig. S7B). More red dots in 2D-tsne plots (Suppl. Fig. S7C) of M1 + NextA macrophages indicate increased secretion of proinflammatory cytokines IL-18 and GM-CSF with NextA treatment compared to M0 and M1 macrophages. However, IL-12p70 was not affected by HDAC6 inhibition. The signal intensity scatter plots indicate the level of expression of respective cytokines shown in 2D-Tsne plots Suppl. Fig. S7E. On the other hand, secretion of immunoregulatory cytokine IL-13 and soluble CD40L by M2 macrophages was not affected by NextA, whereas growth factor VEGF was significantly decreased with HDAC6 inhibition (Suppl. Fig. S7D). The signal intensities of corresponding cytokines from each cell are shown as scatter plots in (Suppl. Figs. S7E-F). Overall, inhibition of HDAC6 with NextA enhanced the inflammatory function of human M1 macrophages but also decreased the M2 function similar to murine BMDMs.
We applied a macrophage ACT approach similar to SM1 murine melanoma with human humanized NSG-SGM3 mice harboring BRAF V600E mutated human melanoma tumor xenografts. Before intratumor implantation, macrophages were verified for M1 polarization by qRT-PCR analysis to express M1 markers CD80, CD86, and TNF (Suppl. Fig. S8A). Intratumor ACT of human macrophages into PDX tumors significantly reduced tumor size, mirroring the results from the SM1 melanoma model (Suppl. Fig. S8B). Since we did not observe significant differences in the effect of vehicle-treated M1 macrophages with the SM1 murine melanoma model, we did not include the M1 group in this experimental design. Further analysis of the immune cell composition of the TME revealed an increase in antitumor M1 macrophages as a fraction of total CD45 + cells. Similar to the SM1 model, we observed an increment in the M1/M2 ratio in those tumors subjected to M1 + NextA treatment (Suppl. Fig. S8C). Despite the trends in the number of M1 and M2 macrophages not being significant due to small sample size (n = 5 per group), tumor-associated macrophages’ trends were similar to the SM1 model. However, an important increase in the M1/M2 ratio (p-value, 0.0703) suggested that NextA-treated M1 macrophages provided antitumor immunity. Histological analysis of humanized melanoma tumors was consistent with the flow cytometry data. We observed increased M1 macrophages in the vicinity of blood vessels and towards the periphery, along with scattered distribution within the tumor, suggesting an increased infiltration of proinflammatory M1 macrophages. On the contrary, the number of M2 macrophages within the tumor was substantially lower, similar to the histological analysis of SM1 tumors (Suppl. Fig. S8D). Overall, the NSG-SGM3 model with a humanized immune system recapitulated the observations in SM1 murine melanoma model supporting HDAC6 inhibitor-treated M1 macrophages as a potential cell therapy to treat solid tumors.








Add Comment